Overview of Our Plant Systems Biology Research
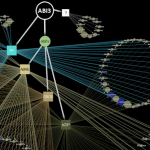
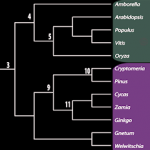
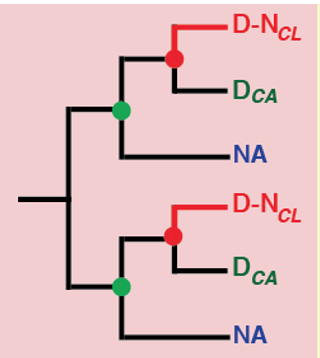
The over-arching focus of our research has been to develop Plant Systems Biology approaches to predictively model how internal and external perturbations affect processes, pathways & networks controlling plant growth, development and evolution. We apply these approaches to study the Systems Biology of Nitrogen Use Efficiency (NUE) which have uncovered the regulatory networks coordinating a plants response to sensing nitrogen sources in its environment and internally. These studies have identified the N-regulatory networks and key hubs that coordinate nitrogen regulation of metabolic processes (N-assimilation), cellular processes (circadian rhythm) and developmental processes (N-foraging in roots), providing a holistic or systems view of NUE. We embodied the tools we developed for regulatory network analysis, and other visualization/analysis tools into a software platform called VirtualPlant (www.virtualplant.org) to broadly enable Systems Biology approaches across the plant community. We have also developed approaches and tools to enable comparative genomic studies across all seed plants, which includes the creation of “BigPlantv1.0” – ( the result of an automated pipeline for orthology identification and construction of phylogenomic-scale trees – currently composed of 22,833 sets of orthologs from the genomes of 150 plant species. The BigPlant phylogenomic framework enables researchers to identify overrepresented functional gene categories at major nodes in seed plant phylogeny, revealing critical genes and biological processes important to Angiosperm evolution.
Our Plant Systems Biology Studies focus on Four main areas:
• Nitrogen regulatory networks:The Systems Biology of Nitrogen use efficiency (NUE)
• VirtualPlant: A software platform for Plant Systems Biology
• Evolutionary Genomics:A functional phylogenomic approach to seed plant evolution.
• EVONET: A Phylogenomic and Systems Biology approach to identify genes underlying plant survival
in marginal, low-Nitrogen soils
Funding: Our research is supported by NIH, NSF, DOE, and Zegar Foundation



![]()